Class 12 Biology Chapter 6 Molecular Basis of Inheritance The answer to each chapter is provided in the list so that you can easily browse through different chapters Assam Board HS 2nd Year Biology Chapter 6 Molecular Basis of Inheritance Question Answer.
Class 12 Biology Chapter 6 Molecular Basis of Inheritance
Also, you can read the SCERT book online in these sections Solutions by Expert Teachers as per SCERT (CBSE) Book guidelines. These solutions are part of SCERT All Subject Solutions. Here we have given Assam Board Class 12 Biology Chapter 6 Molecular Basis of Inheritance Solutions for All Subjects, You can practice these here.
Q.10. What is meant by semiconservative of DNA replication?
Ans : According to semi consecutive hypothesis of DNA replication the unzipping of DNA strands takes place and complementary strand of each unzipped strand in formed is such way that the daughter strands are exact replica of the parent DNA strands.
Q.11. What is Frame shift insertion?
Ans : Deletion or insertion of one or a few nucleotides in the DNA molecule results mutation. Therefore, such mutation that results in shifting of reading frame backward or forward by one or more nucleotide is called frame shift mutation.
Consider a statement that is made by following words :
RAM HAS RED CAP
If a letter B is inserted between HAS and RED then the statement would be –
RAM HAS BRE DCA P
If it is followed by word ‘BI’ and ‘BIG’ then the statements would be:
RAM HAS BIR EDC AP
RAM AIAS BIG RED CAP
Same can be repeated by deleting R, E and D one by one as follows:
RAM HAS RED CAP
RAM HAS EDC AP
RAM HAS DCA P
RAM HAS CAP
Q.12. What are the functions of DNA polymerase.
Ans : In DNA replication a set of enzymes are involved. Of these the main ‘enzyme is referred to as DNA polymerase. It catalyses polymerization of DNA nucleotides of a large number in a very short time. The time required to polymerize depend, upon the number of base pair involved. Generally the speed of polymerization is 2000 base pairs per second. A great amount energy is required for polymerization process. Deoxyribonucleotide triphos phate not only provide energy but also acts as substrate.
Due to requirement of high amount energy for separation of two strands of DNA molecules during replication, a small fork called replication fork occurs in the DNA helix. During replication, the DNA-polymerase catalyse polymerization in one direction, 5′-3′ in one strand and in other the direction is 3′-5′. The polymerization in 3′-5′ direction is continuous while in 5′ – 4 3′ direction it is discontinuous. The discontinuous fragments are later joined by enzyme DNA ligase.
The DNA replication does not occur spontaneously or does not occur at any place. The origin of replication at particular site is determined by the requirement of a particular piece of DNA for the purpose of recombination. Generally a vector locates the origin of replication. In eukaryotic cells, DNA replication takes place in S-phase of cell-cycle.
Q.13. What are the functions of mRNA and tRNA.
Ans : (a) Messenger RNA (mRNA) : It acts as messenger to carry the message of the genetic code from the DNA of the nucleus to the site of protein synthesis, the ribosomes. It is also known as nuclear RNA as it is found only in the nucleus. It acts as a template for the translation of the DNA code in the formation of specific protein. It is formed as a complementary strand to one of the two strands of DNA which acts as a template. So, mRNA molecule possesses the same sequence of nitrogenous bases as is found in the template DNA. But at the time of synthesis of MRNA, the thymine of DNA is replaced by uracil.
(b) Transfer RNA : Transfer RNA or TRNA is the second most common RNA of a cell. It is the smallest molecules of RNA. tRNA has great affinity for specific amino acids and ATP. It brings activated amino acids to the proper site on the mRNA template. This RNA is synthesized in the particular region of DNA molecule and has complementary base sequence as in DNA.
In addition to the above three types of RNA, there is another type of RNA which is found in virus and so known as viral RNA or VRNA. This is found in TMV, influenza virus, tobacco necrosis virus.
Q.14. Elucidated the lac operon model of gene expression regulation.
Ans : The control of protein synthesis has been elucidated in E. coli by Francis Jacob and Jacques Monod at the Pasteur Institute in Paris in 1961. It was found that when sugar lactose is added to the cultures of E. coli, it induces the synthesis of three enzymes required for degradation of lactose to glucose and galactose. Enzymes are β galactosidase, permease and transacetylase. Process is referred as induction and enzymes formed are called inducible enzymes. The genes for these three enzymes lie close to each other and are linked.
These genes are called structural genes as they carry necessary information to code for amino acid sequence and thus finalise the structure and function of enzymes (proteins) of the pathway. The lactose or lac operon of E. coli has three structural genes and its tryptophan operon has 55 such genes. These genes are regulated as a unit by a single switch called operator. Promoter gene marks the site at which transcription of mRNA starts and where RNA polymerase enzyme birds. The entire unit is called operon.
The action of structural gene is regulated by operator site with the help of a repressor protein produced by the action of gene “i” called as regulator gene. The genes are expressed or not expressed depends upon whether the operative switch in on or off. With operator switch on, the three genes are transcribed by RNA polymerase into a single stretch of mRNA covering all the three genes. Each gene segment is called cistron and long m RNA covering all the cistrons is known as polycistronic. If lactose is removed from the medium, the enzymes needed for degradation are not produced.
The repressor substance: may combine with operator gene to repress its action in two ways.
| Sl. No. | CONTENTS |
| Chapter 1 | Reproduction in Organisms |
| Chapter 2 | Sexual Reproduction in Flowering Plants |
| Chapter 3 | Human Reproduction |
| Chapter 4 | Reproductive Health |
| Chapter 5 | Principles of Inheritance and Variation |
| Chapter 6 | Molecular Basis of Inheritance |
| Chapter 7 | Evolution |
| Chapter 8 | Human Health and Disease |
| Chapter 9 | Strategies for Enhancement in Food Production |
| Chapter 10 | Microbes in Human Welfare |
| Chapter 11 | Biotechnology: Principles And Processes |
| Chapter 12 | Biotechnology and its Applications |
| Chapter 13 | Organisms and Populations |
| Chapter 14 | Ecosystem |
| Chapter 15 | Biodiversity and Conservation |
| Chapter 16 | Bioresources of Assam |
| Chapter 17 | Environmental Issues |
(I) IAC OPERON (INDUCIBLE OPERON) :
In the operon is generally off, as a result, there is no transcription and thus no formation of proteins (enzymes)
However, it can be turned on if a metabolite is provided to the bacterium from outside. The consists of three structural genes, Lac Z, Lac Y and Lac A. The added metabolite comes in contact with active repressor bound to operator, leads to change in structure of repressor and repressor is removed from operator. Operon gets transcribed and enzymes are produced. The process continues till metabolite (inducer) is consumed. After the consumption of inducer (metabolite) repressor again gets its tertiary form, binds it to the operator and switch off the operon.
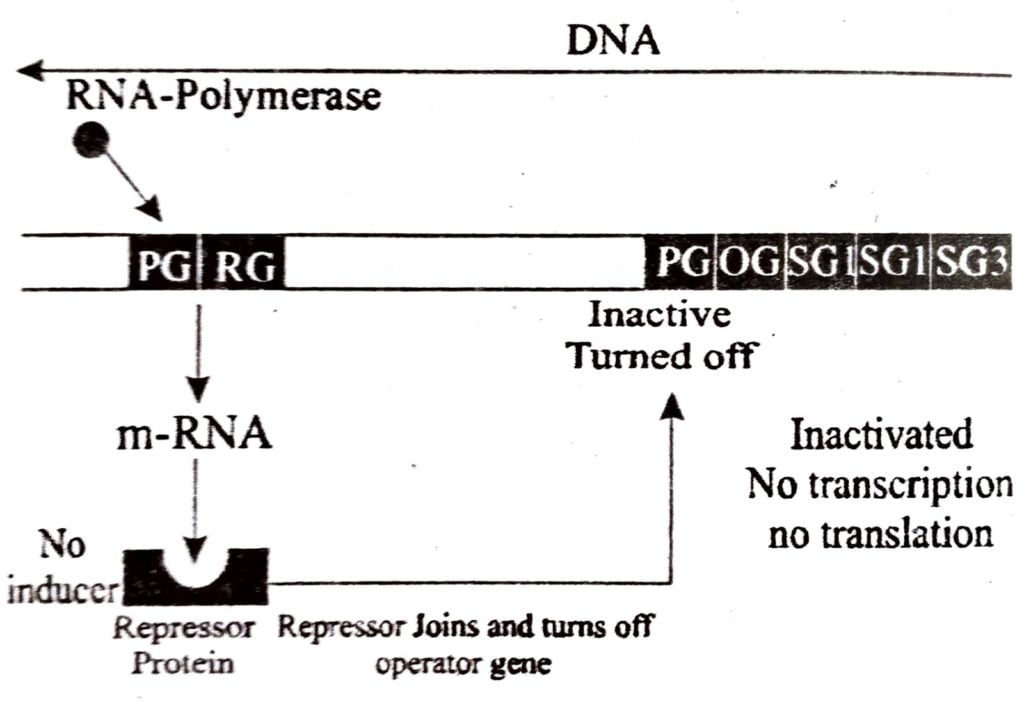
In Lac operon lactose when added enters the cells by the action of enzyme permease few molecules of which are usually present in cell. Lactose binds itself to active repressor leading to change in its structure. As a result repressor now fails to bind itself to the operator. Now RNA polymerase starts the process of transcription of operon by binding to promoter site P. All the three enzymes are formed viz. β galactosidase, permease and transacetylase. Finally, all lactose molecules are used up. Now inactive repressor turns active, attaches itself to operator and finally switches off the operon.
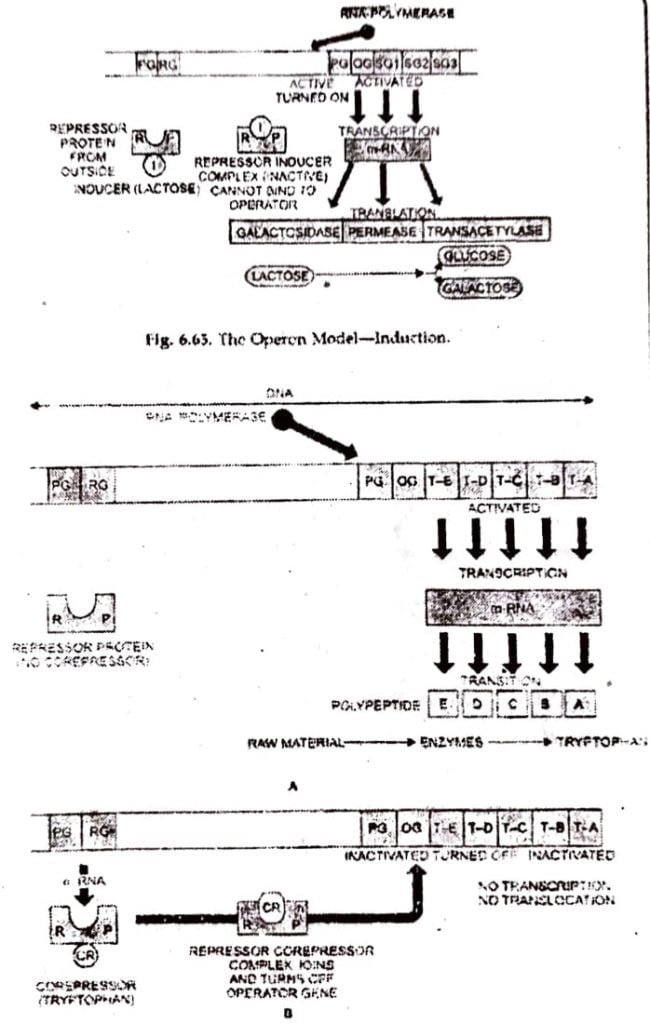
(II) REPRESSIBLE OPERON (TRYPTOPHAN OPERON AND ARGININE OPERON)
In this system, operon is usually on so that transcription occurs normally to synthesize enzymes But operon can be switched off due to non- requirement of metabolite.
In tryptophan operon, formation of tryptophan (an amino acid) needs action of five enzymes in succession. The formation of tryptophan is on because regulator gene R forms an inactive repressor called aporepressor which does not attach itself to operator site. As operator site is free of repressor, the operon system remains on and all the five enzymes needed for tryptophan formation are synthesized.
When tryptophan accumulates or added, few molecules of tryptophan act as co-repressor and binds to inactive repressor (repressor-corepressor complex) which turns active and attaches itself to operator, thus switching off the operon.
Q.15. What is DNA finger printing? Mention its application.
Ans : Technique of using DNA fragments, resulting from restriction endonuclease enzyme cleavage to identify particular individuals, is called DNA finger printing.
Introduction: DNA finger printing technique was developed by Alec Jffery (1985, 86) at Leicester University, United Kingdom, Inheritance of
DNA is very stable. Every person has specific pattern of DNA sequence which shows a combination of DNA sequence of both mother and father. Study of DNA finger prints helps in establishment of identity of a person, identification of criminals in case of a murder or rape, and paternity test in case of disputed parentage. This technique helps in identification even when the stains on victims clothes etc. are even several years old.
In India, first test of DNA finger printing was done in June, 1989 for setting a disputed parentage in Madras. Laboratory for DNA finger printing is situated in Hyderabad at the Centre for Cell and Molecular Biology (CCMB). Paternity dispute cases are much more common in India and most of them are referred to CCMB. for DNA evidence.
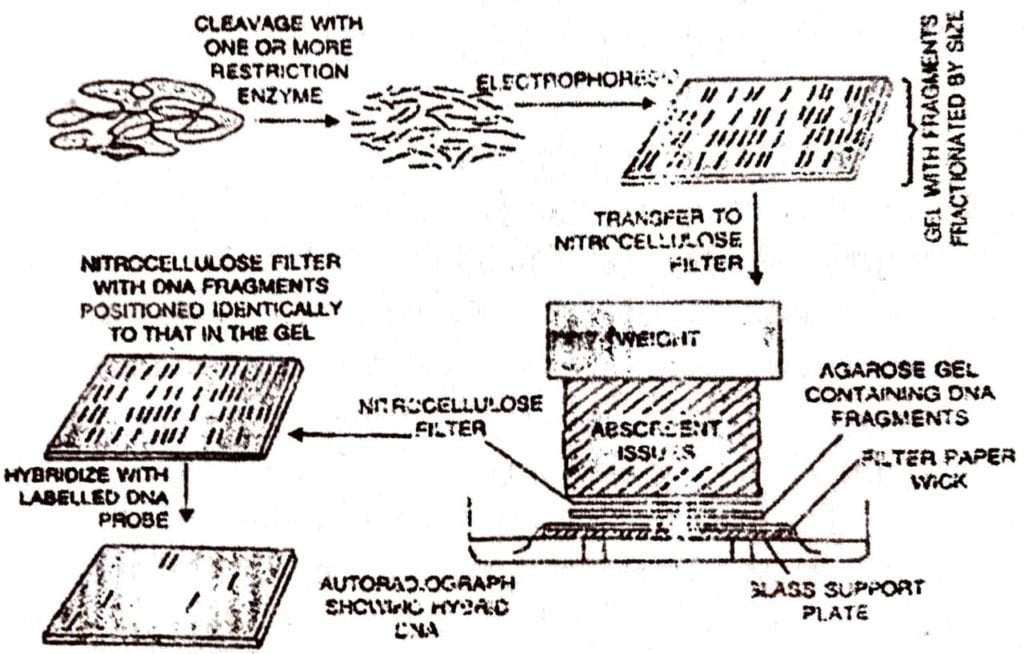
Southern Blotting technique of DNA finger printing:
VNTR (Variable number of tandem repeats) technique involved Southern blot hybridisation using radiolabeled VNTR as a probe. The steps were:
(a) For DNA finger printing, DNA is isolated from any body cell or (n) even from blood stains, semen stains or hair roots.
(b) DNA sample is first digested with a restriction endonuclease enzyme and the digested sample is then gel electrophoresed.
(c) DNA bonds in the gel are denatured into single strands by treating them with alkali solution.
(d) Gel is laid on the top of a buffer saturated filter paper, placed on a (p) solid support (e.g.glass plate), with its two edges immersed in the buffer.
(e) A sheet of nitrocellulose membrane is placed on top of the gel and a stack of paper towels is placed on the top of this membrane (acts as an absorbent tissue).
(f) A weight of about 0.5 kg is placed on the top of pack of paper towels. Leave this arrangement for a few hours or over night.
The buffer solution is drawn up by the filter paper wick, and passes through the gel to the nitrocellulose membrane and finally to the paper towels. While passing through the gel, the buffer carries with it single stranded DNA molecules, which bind to the nitrocellulose membrane, while the buffer passes through it to the paper towels.
(g) Next day, paper towels are removed and discarded.
(h) Nitrocellulose membrane with single stranded DNA bands is incubated at 80°C for the 2-3 hours to fix the DNA bands permanently on the membrane.
(i) This membrane now has a replica of DNA bands from agarose gel and can be used for hybridization with specific labelled DNA or RNA probes.
(j) The membrane may then be washed to remove any unwound DNA and X-ray film is exposed to the hybridized membrane to get autoradiograph.
Significance: DNA finger printing shows polymorphism of DNA which is used for identification of a .. with much more certainty that has been possible through techniques of blood groups since the number of blood groups available becomes a limitation. The technique has shown the possibility of two persons having same pattern of DNA finger prints is very remote (exception monozygotic i.e. identical twin)

Hi, I’m Dev Kirtonia, Founder & CEO of Dev Library. A website that provides all SCERT, NCERT 3 to 12, and BA, B.com, B.Sc, and Computer Science with Post Graduate Notes & Suggestions, Novel, eBooks, Biography, Quotes, Study Materials, and more.


