Class 12 Biology Chapter 11 Biotechnology: Principles and Processes Solutions to each chapter is provided in the list so that you can easily browse through different chapters Assam Board HS 2nd Year Biology Chapter 11 Biotechnology: Principles and Processes Question Answer.
Class 12 Biology Chapter 11 Biotechnology: Principles and Processes
Also, you can read the AHSEC book online in these sections Solutions by Expert Teachers as per AHSEC (CBSE) Book guidelines. These solutions are part of AHSEC All Subject Solutions. Here we have given Assam Board Class 12 Biology Chapter 11 Biotechnology: Principles and Processes Solutions for All Subjects, You can practice these here.
Biotechnology: Principles and Processes
Chapter – 11
BIOTECHNOLOGY
Very Short Answer Type Questions
Q.1. What do you understand by PCR?
Ans : PCR means Polymerase Chain Reaction. With the help of this one can multiply any fraction of DNA.
Q.2. What is Biotechnology?
Ans : Technical use of the knowledge of biology for production of useful substances is called biotechnology.
Q.3. Define recombinant DNA technology. What is the popular terminology of it.
Ans : Reconstruction of DNA by addition or deletion of genes is called DNA recombinant technology.
Q.4. What do you mean by bioreactors?
Ans : Bioreactor is a kind of formentor which provides a control environment for growth of microorganism for obtaining a desired product.
Q.5. What is cloning vector?
Ans : In DNA recombinant technology gene from one DNA is remove and transferred to another DNA. This is called gene cloning. For this purpose a vector is necessary to carry the DNA fragment to the target DNA. Generally some viruses and plasmids are used as vector for this purpose.
Q.6. Expand the terms Ori and EcoRI.
Ans : Ori is a plasmid vector. It means origin of replication. For all replication purpose this is essential. EcoRI stands for Escherichia Coli Ry 13. It is also a plasmid vector.
Q.7. What is gene gun?
Ans : During recombination DNA gene collected from other source is inserted in various ways, the most common method is use of plasmid as vector. There another method by which gene is pushed into cell by force. The device acts like gunshot and hence it is called gene gun.
Q.8. What is the function of DNA ligase?
Ans : It is an enzyme used to join two pieces of DNA.
Q.9. In respect to mode of action how can you differentiate enzyme exonuclease from endonuclease?
Ans : Endonuclease cut DNA to separate nucleotide sequence away from the ends. Exonuclease cut nucleotide sequence from the ends of the DNA.
Q.10. What is competent cell?
Ans : The cells produced by applying biotechnology in called competent cells.
Q.11. Who first generated the recombinant DNA molecules?
Ans : Stanley Cohen and Herbert Boyer.
Q.12. What is gene cloning?
Ans : Gene cloning means isolation of gene and then making multiple cOpies of it by inserting it into a bacterial cell or another organisms.
(B). Fill up the blanks:
Q.1. Each restriction endonuclease recognizes a specific ____ in the DNA.
Ans : Palindromic sequence.
Q.2. In plants recombinant DNA is injected into the nucleus by a method known as _____
Ans : Gene gun.
Q.3. Ri and Ti plasmids are found in ____
Ans : Agrobacterium.
Q.4. PCR stand for ____
Ans : Polymerase.
Q.5. ____ are known as molecular scissors.
Ans : Endonuclease.
Q.6. The first restriction endonuclease recognized is____
Ans : Hind III.
Q.7. ____ deals with large scale production and marketing of products and process using live organisms, cells or enzymes.
Ans : Biotechnology.
Q.8. Most commonly used bioreactors are of _____
Ans : Membrane bioreactor.
Q.9. A piece of DNA is introduced in a host bacterium through ____
Ans : Plasmid.
Q.10. Gel electrophoresis is used for _____
Ans : Separate DNA fragments.
Q.11. ____ a crown gall bacterium is called natural generic engineer of plants.
Ans : Agrobacterium tumefaciens.
Q.12. Genetic engineering is also known as ____ DNA technology.
Ans : Recombinant.
Q.13. Large scale production of biotechnological products involve use of
Ans : Cloning.
(C). Select the true and false statements.
Q.1. Bacteria protect their, nucleoid from restriction endonuclease through methylation.
Ans : True.
Q.2. Transgenic plants are developed by introduction of foreign genes.
Ans : True.
Q.3. Function of restriction enzyme is to cut DNA at the ends.
Ans : False.
Q.4. DNA fragments are joined by enzyme polymerase.
Ans : False.
Q.5. Bacteria protect themselves from virus by fragmenting viral DNA with endonuclease.
Ans : True.
Q.6. Restriction enzymes are also known as molecular markers.
Ans : False.
Q.7. An antibiotic resistance gene in a vector usually helps in the selection of transformed cells.
Ans : True.
Q.8. The most common bacterium used in genetic engineering in Escherichia.
Ans : True.
Q.9. Hargovind Khurana was awarded the Noble Prize for the development of PCR technique.
Ans : False.
Q.10. Genetic engineering is a technique which involves deliberate manipulation of genes within or between the species.
Ans: True.
Q.11. Plasmids are circular extra chromosomal DNA in Bacteria.
Ans : True.
Q.12. DNA ligases are used to cleave a DNA molecule.
Ans : False.
Q.13. Bacteria protect them from viruses by fragmenting viral DNA with endonuclease.
Ans : True.
II. Short Questions 2 Marks :
Q.1. What DNA ligase?
Ans : It is an enzyme DNA. It catalyzes the linkage between two free ends of double stranded DNA chains by forming a phosphodiester bond between them, as in the repair of damaged DNA.
Q.2. Define genetic engineering. Mention the tools used in genetic engineering.
Ans : Genetic engineering can be defined as the science dealing with the correction of nature’s mistake at the level of hereditary unit, the DNA and to improve the quality of organism for purposes by implanting desired genes.
Tools of genetic engineering :
(i) Enzyme lysozyme is used to dissolve cell wall of bacteria to bring () out DNA fragments.
(ii) Enzymes restriction endonuclease and exonuclease which is used (1) to cut the desired segments of DNA for the purpose of recombination.
(iii) Enzyme ligase used to join two cut ends of DNA fragments.
(iv) Enzyme DNA polymerase is used to synthesise complementary DNA strand on DNA template.
(v) PCR is used to multiply DNA fragments.
(vi) Vectors : Plasmid and viruses are used to transport the DNA fragments inside the cell.
Q.3. What is gene gun? Give its working principles.
Ans : During recombination DNA gene collected from other source is inserted in various ways, the most common method is use of plasınid as vector. There is another method by which gene is pushed into cell by force. The device acts like gunshot and hence it is called gene gun.
Q.4. What are transgenic plants? Give examples.
Ans : The plants which have been modified used gene from other species are called transgenic plants. The other species may be any microorganism, animal or distantly related plants.
Bt, cotton is a transgenic plant. Cotton cultivation is affected all over world by budworm that destroys the buds. The toxin produced by a bacteria called Bacillus thuringiensis is found to be inhibitory to such worms. Using genetic engineering, the gene responsible for coding the toxin in the bacteria had been isolated and inserted into the plant genome. This has provided the cotton plant resistance to budworms.
Q.5. How is DNA isolated in purified form from a bacterial cell?
Ans : DNA is isolated from bacteria using restriction endonuclease it is then cleaned and each piece is separated using gel electrophoresis to make each piece free from cell debris. The fragments of DNA then multiplied using PCR.
Q.6. What is recombinant DNA? How do enzymes restriction endonuclease and DNA ligase help its formation?
Ans : Genetically engineered DNA prepared by transplanting or splicing genes from one species into the cells of a host organism of a different species is called recombinant DNA. Such DNA becomes part of the host’s genetic makeup and is replicated.
The enzyme endonuclease is used to cut the required piece of DNA in an organism. This enzyme can identify the particular sequence of nucleotide where it should cut. The cut piece is the brought out by a special technique. The cut piece of DNA is then pushed into the cell of the target organism using suitable vector or gene gun. Ligase enzyme is then used to join the cut piece with the DNA of the host.
Q.7. Name the source organism from which T₁ plasmid is isolated. Explain the use of this plasmid in biotechnology.
Ans : T₁ plasmid is obtained from Agrobacterium tumefaciens. This plasmid is used as vector for carrying the required DNA fragment inside the cell of the host. This is widely used for DNA recombination particularly in developing transgenic plants.
Q.8. A recombinant DNA is formed when sticky ends of vector DNA and foreign DNA join. Explain how the sticky ends are formed and get joined.
Ans : Endonuclease cut at the both ends of palindrome sequences of the DNA in such way that sticky ends form in both ends after removal of the cut ends join easily. Such cut DNA fragments will be in the sequence of 5 → 3 and 3’→ 5′. Both these ends then can easily join.
Q.9. What are selectable markers? What is their use in genetic engineering?
Ans : Selectable markers are some genes such as antibiotic resistant genes. These genes are inserted into the cloned cells. This is to test whether the cloned cell has recombinant DNA. Because in many cases the apparently cloned cell may not have the desired r combined DNA. To test whether the cloned cell are really cloned or not the selectable marker genes are infused into the cells. The cells containing antibiotic resistant selectable marker are then cultured in medium containing that particular antibiotic. In the medium only those cells will survive which have selectable markers and others will dye. Those that survive are really contain the desired recombinant DNA.
Q.10. Why is Agrobacterium mediated genetic transformation described as natural genetic engineering in plants?
Ans : Agrobacterium is the only organism known which has developed technique of gene transfer from it to some plants and thereby cause an infection in the plant called ‘gall’. It has developed this technique in course of evolution. Through such transfer of its DNA to the plant it passes message to the plant to prepare appropriate nutrition for it. Such gene transfer involve complicated mechanism which is equivalent to complex process of genetic engineering. Therefore naturally it is called natural genetic engineering. Actually scientists have learned the technique of recombinant DNA technology from Agrobacterium.
Q.11. Do eukaryotic cells have restriction endonuclease? Justify your answer.
Ans : Restriction enzymes are a class of enzymes called endonucleases. Endonucleases are able to cut in the middle of the DNA backbone or the phosphodiester bonds. A different class of enzymes called exonucleases cut the DNA backbone, but only from the ends-either from the 3′ end or the 5′ end. Most restriction endonucleases are prokaryotic-in origin. However, there are several found in eukaryotic cells, including our own. In eukaryotes they are not referred to as restriction enzymes, just endonucleases. An example of an endonuclease in eukaryotes is Apn 1, isolated from yeast. This enzyme helps prevent DNA damage from environmental agents.
Q.12. While doing a PCR, denaturation step is missed, what will be its effect on the process.
Ans: There are 6 steps in PCR process. Of these the denaturation of DNA is the first step. If this step is missed the next step. Annealing step will not follow and thereby subsequent steps will also not follow and ultimately amplification of DNA will not be accomplished.
Q.13. Why enzyme cellulose is used for isolating genetic material from plant cells but not animal cells?
Ans : The enzyme cellulose is used to separate the genetic material from the plant cells this enzyme can act only on plants but not the animals. There is no role is played by enzyme cellulose on animal body. They can break the cell wall and content will released.
Q.14. What are major steps involved in gene cloning technique?
Ans : Gene cloning technique involves the following steps:
(i) Isolation of required DNA, generally called donor DNA out of a suitable organism by biochemical technique.
(ii) Breaking of DNA into fragments using restriction enzymes.
(iii) Isolation of suitable bacterial plasmid and cutting it at a place by restriction enzyme and joining the donor DNA fragment with it with the help of an enzyme called ligase. The plasmid containing parent DNA and donor DNA.is called recombinant DNA.
(iv) The recombinant DNA is then inserted within similar bacteria through cloning.
(v) Culturing the cloned bacteria containing fragment of donor DNA.
(vi) Selection of appropriate bacterial colony which has incorporated Sil the sought after DNA fragment. For example, the different segments of DNA inserted in plasmid may cause bacteria to synthesize different substances. It is possible therefore to create bacteria acquiring ability to synthesize any substance like insulin, haemoglobin, interferon etc.
Q.15. What are molecular scissors? Give at least two examples of it?
Ans : Restriction enzymes are called the molecular scissors The example of restriction enzymes are – EcoRV and Eco RT.
III. Short Questions 3 Marks :
Q.1. State the basic steps involved in genetically modifying an organism.
Ans : The following basic steps are involved in genetic modification of organisms.
(i) Isolation of DNA.
(ii) Fragmentation of DNA.
(iii) Isolation of desired fragments of DNA.
(iv) Joining of DNA fragments into vector.
(v) Transferring recombined DNA into host.
(vi) Culturing of host cells for multiplication of recombined DNA.
Q.2. Name the specific enzymes used to break the cells of organisms like bacteria, eukaryotic płant and fungus and release DNA along with other macromolecules.
Ans :
| Aganisms | Enzymes |
| Bacteria | Lysozyme |
| Plant cells | Cellulase |
| Fungi | Chitinase |
3. Distinguish between :
(i) Plasmid DNA and Chromosomal DNA.
Ans : (a) Plasmid is the circular DNA while chromosomes are found in higher groups of organism as carrier of heredity.
(b) Plasmid is found in bacteria and plays great role in genetic engineering.
(ii) Exonuclease and Endonuclease.
Ans : Exonuclease is an enzyme used to cut the DNA for the purpose of cloning while the endonuclease is termed as bio scissors. Both the enzymes are very useful in gene cloning.
Q.4. What is meant by recognition sequence? Write the recognition site of EcoRI and where it cuts the DNA.
Ans : Endonuclease cut DNA at specific point to separate certain sequence of base pair nucleotides. It therefore must recognise the Palindromic nucleotide sequence in DNA before cutting. The sequences are either 5 → 3 or 3 → 5 and may contain a few to they base pairs.
There are 3 types of endonucleases – Type I, Type II and Type III which cut sequence at different base pairs. Of these the types II remarkable for their palindrome. They are also very stable. In palindrome, the base sequence such that if read from both end they appear to be same.
5′ GAA ↑ AAG 3′ 5° GAA ↑ AAG 3′
Single Stand DNA Double Stranded DNA
(If read from right to left and left to right it will be the same sequence in both strands)
In palindrome with rotational symmetry the base sequence in the first half of one strand DNA double helix is the mirror image of the second half of its complementary stand. Thus the base sequence in both the strands read the same when read from 5′- 3′ end and 3′-5′ end both sands. Thus recognition sequence of ECORI are
5′ →3′ or 3 → 5 the
5′ GAA | TTC 3′
3′ CTT | AAG 5″
ECORI recognition site
Q.5. What is downstream processing in respect of biotechnology?
Ans : Any product made through biotechnology is not a finished product. It may have impurity, it may contain harmful by products also. So these need to be refined. The processes which purifies materials as finished products is called downstream process. Further, preservation and storing also need to be addressed appropriately. In case of pharmaceuticals clinical trials need to be carried out. The quality control measure is another area of downstream process.
Q.6. Write the major steps involved in gene cloning.
Ans : Genes are specific parts of DNA. For cloning specific gene, it must be separated from the DNA and then it should be attached with a vector like plasmid or bacteriophage DNA (vector are carriers). As the plasmid or the bacteriophage DNA multiplies within the bacteria or the bacteriophages the gene inserted in them also multiplies. For this purpose, the following methods are applied.
(i) At first the plasmids DNA found in bacteria are brought out of the 6. bacterial cell by lysing the bacterial cell. The plasmid which is used to carry the gene to be cloned is also called passenger or vehicle DNA or Vector. In place plasmid, bacteriophage DNA can also be isolated and used as vector.
(ii) In the next step, the gene to be isolated and cloned is identified in the DNA of another bacteria or organism and is bought out.
(iii) Now using an enzyme called restriction endonuclease the specific part of the DNA carrying the specific gene is cut out. With the same enzyme the specific part of the plasmid where the gene will fit is opened by cut and the equivalent part of the plasmid is taken out so that the inserted gene fits will in the plasmid. The cut end of the plasm and the DNA well have sticky ends in one of their single strand. These sticky ends help them to join with their complementary parts.
Q.7. Match the items of column A with the items of column B.
| Column “A” | Column “B” |
| (i) Genetic engineering | (a) Vector + insert |
| (ii) Sticky ends | (b) Restriction Endonucleases |
| (iii) Electroporation | (c) Large scale production |
| (iv) Vehicle DNA | (d) Vector less gene transfer |
| (v) r DNA | (e) Recombinant DNA technology |
| (vi) Bioreactors | (f) Cloning Vector |
Ans :
| Column “A” | Column “B” |
| (i) Genetic engineering | (e) Recombinant DNA technology |
| (ii) Sticky ends | (b) Restriction Endonucleases |
| (iii) Electroporation | (d) Vector less gene transfer |
| (iv) Vehicle DNA | (f) Cloning Vector |
| (v) r DNA | (a) Vector + insert |
| (vi) Bioreactors | (c) Large scale production |
Q.8. Give reasons of recombinant DNA formation from two sources of DNA.
Ans : DNA recombination is done to add certain quality gene to the DNA by deleting some part of a DNA and adding another piece there taken from DNA of another organism. The inserted DNA piece must have specific nucleotide sequence that code for certain quality which is not thee in the host DNA. It is therefore a recombinant DNA carries DNA from host and doner.
Q.9. State the principle underlying ‘gel electrophoresis’ and mention two applications of this technique in Biotechnology.
Ans : The device uses a substance called agarose gel. Any substance having twɔ different fractions will move opposite to each other depending upon the electric charge. Some will move towards positive end and another will move towards negative end and thus both can be separated. This device is extensively used for separation of
(i) DNA fraction done by endonuclease. The lighter fragment will move faster than heavier fragments and thus these can be collected separately.
(ii) It is also extensively used in DNA fingerprinting in which various fractions of DNA are separated by this method.
Q.10. Explain any three methods to force ‘alien’ or recombinant DNA into hosts cells.
Ans : Recombinant DNA is pushed into the host cell by any of the following methods :
(i) Using plasmid vector such as Ti plasmid of Agrobacterium, K12 of E. coli etc.
(ii) Using virus vector such as Lambda, M13 etc.
(iii) Using a special device called gene gun.
Q.11. What is Biotechnology? Discuss the principles of biotechnology.
Ans : The term “biotechnology” was brought into popular usage in mid – 1970s as a result of increased potential for the application of the emerging techniques of molecular biology. Biotechnology is a fast growing applied science. According to definition adopted by European Federation of Biotechnology (1978) “Biotechnology makes it possible, through an integrated application of knowledge and techniques of biochemistry, microbiology, genetics and chemical engineering to draw benefit at the technological level from the properties and capacities of microorganisms and cell cultures.”
OR
The integration of natural science and organisms, cells, parts there of and molecular analogies for products and services is called biotechnology. The term biotechnology was used by Leeds City council (1920) when an institute in biotechnology was set up. We find its applications in even back to 5000 years with the discovery of the production of alcoholic beverages by fermentation. Biotechnology today is undergoing a resurgence in the wide range of applications and the tremendous increasè all over the world. Biotechnology is the application of scientific and engineering principles to the processing of materials by biological agents to provide goods and services.
Principles of biotechnology:
(i) Genetic engineering : The ability to isolate a gene coding for desired product and transfer it to another organism has opened the way either to the more effective production of useful proteins or to the introduction of novel characteristics in the host organism. Thus, this refers to methods to change the chemical structure of genetic material i.e. DNA and RNA and then to introduce into host and alter its phenotype.
(ii) To maintain the microbial contamination free (sterile) ambience in (11). chemical engineering. Due to such type maintenance only desired microbe/ cells will be formed in huge number for manufacture of biotechnological- products like hormones vaccines, blood clotting factors, antibiotics steroids, enzymes etc. It is necessary to have complete aseptic condition.
| Sl. No. | CONTENTS |
| Chapter 1 | Reproduction in Organisms |
| Chapter 2 | Sexual Reproduction in Flowering Plants |
| Chapter 3 | Human Reproduction |
| Chapter 4 | Reproductive Health |
| Chapter 5 | Principles of Inheritance and Variation |
| Chapter 6 | Molecular Basis of Inheritance |
| Chapter 7 | Evolution |
| Chapter 8 | Human Health and Disease |
| Chapter 9 | Strategies for Enhancement in Food Production |
| Chapter 10 | Microbes in Human Welfare |
| Chapter 11 | Biotechnology: Principles And Processes |
| Chapter 12 | Biotechnology and its Applications |
| Chapter 13 | Organisms and Populations |
| Chapter 14 | Ecosystem |
| Chapter 15 | Biodiversity and Conservation |
| Chapter 16 | Bioresources of Assam |
| Chapter 17 | Environmental Issues |
Q.12. Expand the term PCR. Write the principle of PCR.
Ans : PCR technique was developed by Kary Mullis 1985. If one knows the sequence of at least part of a DNA segment to be cloned, a number or copies of that DNA segment can be hugely amplified using polymerase chain reaction. It is able to generate microgram quantities of DNA copies (up to billion copies) of desired DNA segment, present even as a single copy with short time.
The technique is based on principle that when a DNA molecule is subjected to high temperature due to denaturation the two DNA strands separate. As a result two single stranded DNA molecules appear.
Q.13. Explain the process by which a bacterial cell can be made ‘competent’ in recombinant DNA technology.
Ans : The ligated plasmid mixture is introduced into the bacterial cells where they takeup DNA through transformation process. Transformation iş generally carried out by placing activity glowing cells of a bacterium in cold, dilute solution of CaCIP which enhances the ability of bacterial cells to take up foreign DNA. In majority of the cases, E coli is the most preferred host because:
(i) Molecular biology of the bacteria is well understood.
(ii) CaC12 treated cells are highly transformable.
(iii) E-coli transcribes and translates most gram + ve and Gram – ve guess except some actinomycetes genes There are many methods to introduce the ligated DNA into recipient competent alls.
IV. Long Questions 5 Marks :
Q.1. EcoRI is used to cut a segment of foreign DNA to form a recombinant DNA. Show with the help of a diagram.
Ans : Endonuclease cut DNA at specific point to separate certain sequence of base pair nucleotides. It therefore must recognise the Palindromic nucleotide sequence in DNA before cutting. The sequence are either 5′ → 3′ or 3′ → 5′ and may contain a few to they base pairs.
There are types of endonucleases – Type I, Type II and Type III which cut sequence at different base pairs. Of these the types II remarkable for their palindrome. They are also very stable. In palindrome, the base sequence such that if read from both end they appear to be same.
5′ GAA ↑ AAG 3′ 5° GAA ↑ AAG 3′
Single Stand DNA 3′ CTT | TTC 5′
Double Stranded DNA
(If read from right to left and left to right it will be the same sequence in both strands)
In palindrome with rotational symmetry the base sequence in the first half of one strand DNA double helix is the mirror image of the second half of its complementary stand. Thus the base sequence in both the strands read the same when read from 5′- 3′ end and 3′-5′ end both sands. Thus the recognition sequence of ECORI are
5′ → 3′ or 3′ → 5′
5′ GAA | TTC 3′
3′ CTT | AAG 5″
ECORI recognition site
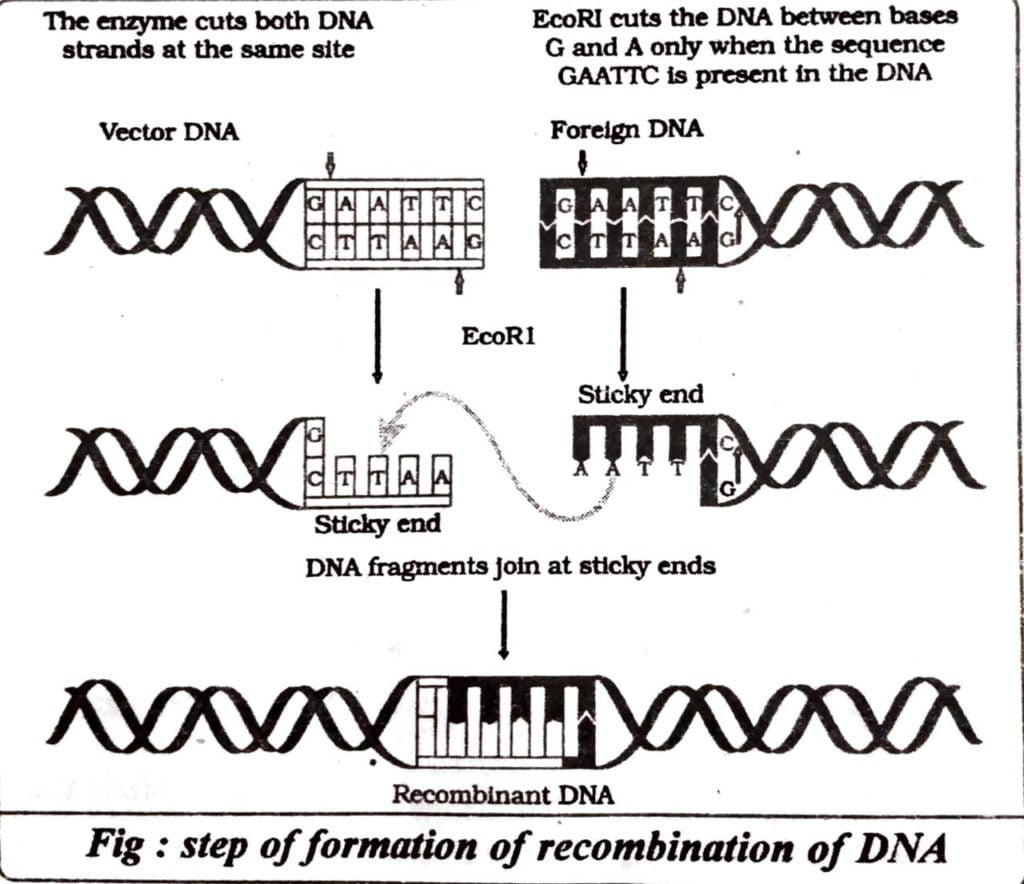
Q.2. What is polymerase chain reaction and how can it be utilized for gene amplification.
Ans : The polymerase chain reaction or PCR multiplies copies of DNA or its segments in vitro using two sets of primers which are small chemically synthesized oligonucleotides complementary to the regions of DNA and an enzyme called DNA polymerase. The enzyme polymerase extend the primers using nucleotides provided in the reaction and the DNA part provided as template. The process of replication can be repeated and a billion copies of the DNA can be multiplied. Thus a small fraction of DNA can be multiplied for study and use in genetic engineering. The following procedure is applied for obtain multiple copies of DNA.
At the start of PCR, the DNA from which a segment is to be amplified, an excess of the two primer molecules, the four deoxyriboside triphosphates and the DNA polymerase are mixed together in the reaction mixture. The following operations are now performed sequentially.
Step 1: The reaction mixture is heated to a temperature (usually 90-98°C) that assures DNA denaturation.
Step 2 : The mixture is now cooled to a temperature (generally 40-60°C) that permits annealing of the primer to the complementary sequences in the DNA; these sequences are located at the 3 ends of the two strands of the desired segment. This step is called annealing.
Step 3 : The temperature is now so adjusted that the DNA polymerase synthesizes the complementary strands by utilizing 3′-OH of the primers; this reaction is the same as that occurs in vivo during replication of the leading strand of a DNA duplex. The primers are extended towards each other so that the DNA segment lying between the two primers is copied.
The completion of step 3 completes the first cycle of amplification; each cycle may take few (ordinarily 1-3) minutes.
Step 4 : The next cycle amplification is separated the newly synthesized DNA strands from the old DNA strands.
Step 5 : Annealing allows the primers to base pair with both the new and did strands, the total number of strands being twice their original number.
Step 6 : Synthesis of new strands takes place, which doubles the number of copies of the desired DNA segment present at the end of step 1. This completes the second cycle.
Thus at each cycle, both new and old strands anneal to the primers and serve as templates for DNA synthesis. As a result, at the end of each cycle, the number of copies of the desired segment becomes twice the number present at the end of the previous cycle. Thus at the end of n cycles 2n copies of the segment are expected; the real values are quite close to this expectation. The cycle may be repeated.
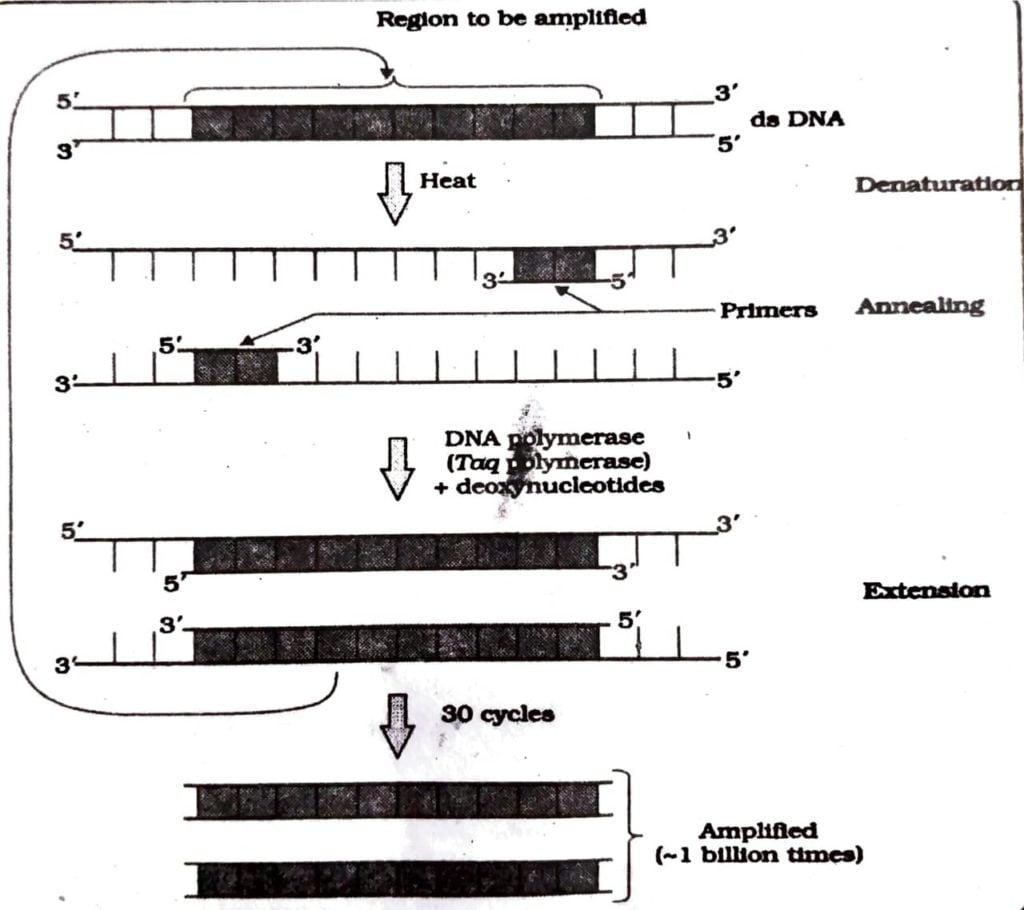
Q.3. What are molecular scissors? Explain their role in the process of recombinant DNA technology.
Ans : Exonuclease and endonuclease are called molecular scissors as they cut the DNA at specific base pair to separate DNA fragments for cloning purposes. These are actually enzymes. Exonuclease cut DNA at the ends that endonuclease cut DNA away from the ends. Before separating the required DNA fragment they recognise the sequence of base pair of nucleotides. As the surgery is done at molecular level these enzyme are called molecular scissors.
Endonuclease cut DNA at specific point to separate certain sequence of base pair nucleotides. It therefore must recognise the Palindromic nucleotide sequence in DNA before cutting. The sequences are either 5′ → 3′ or 3′ → 5′ and may contain a few to they base pairs.
There are 3 types of endonucleases – Type I, Type II and Type III which cut sequence at different base pairs. Of these the types II remarkable for their palindrome. They are also very stable. In palindrome, the base sequence such that if read from both end they appear to be same.
5′ GAA ↑ AAG 3′ 5° GAA ↑ AAG 3′
Single Stand DNA 3′ CTT | TTC 5′
Double Stranded DNA
(If read from right to left and left to right it will be the same sequence in both strands)
In palindrome with rotational symmetry the base sequence in the first half of one strand DNA double helix is the mirror image of the second half of its complementary stand. Thus the base sequence in both the strands read the same when read from 5′ – 3′ end and 3′ – 5′ end both sands. Thus the recognition of ECORI sequence are
5′ → 3′ or 3′ → 5′
5′ GAA | TTC 3′
3′ CTT | AAG 5″
ECORI recognition site
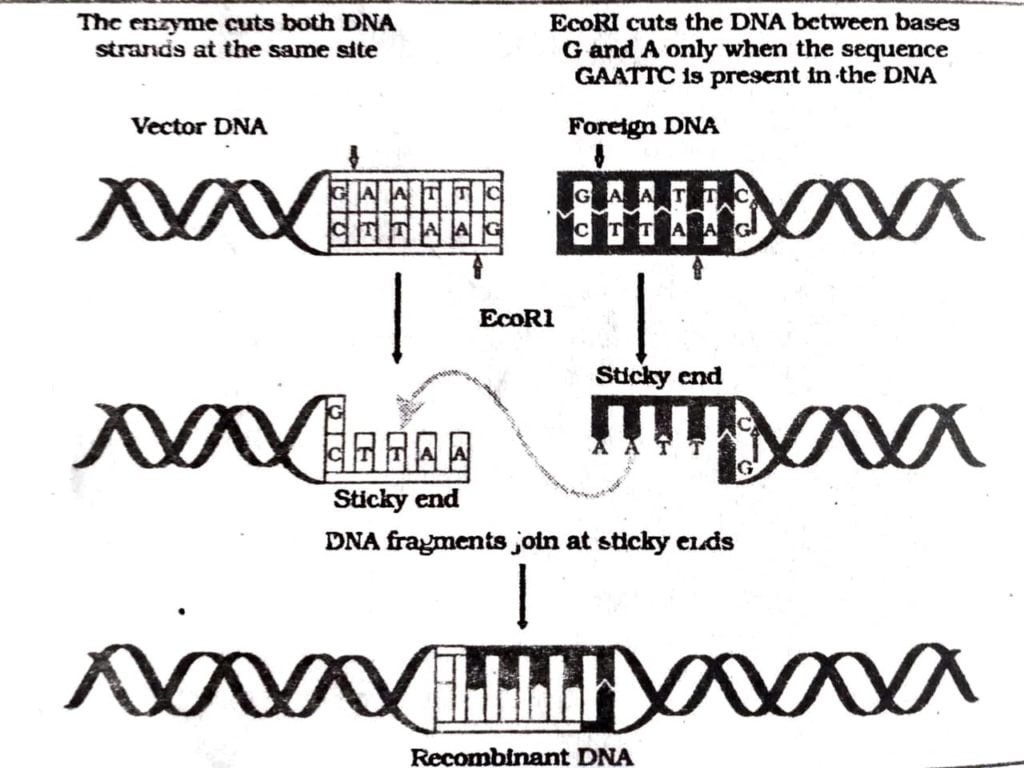
Q.4. Any recombinant DNA with a desired gene is required in billion copies for commercial use. How is the amplification done? Explain.
Ans : The polymerase chain reaction or PCR multiplies copies of DNA or its segments in vitro using two sets of primers which are small chemically synthesized oligonucleotides complementary to the regions of DNA and an enzyme called DNA polymerase. The enzyme polymerase extend the primers using nucleotides provided in the reaction and the DNA part provided as template. The process of replication can be repeated and a billion copies of the DNA can be multiplied. Thus a small fraction of DNA can be multiplied for study and use in genetic engineering. The following procedure is applied for obtain multiple copies of DNA.
At the start of PCR, the DNA from which a segment is to be amplified, an excess of the two primer molecules, the four deoxyriboside triphosphates and the DNA polymerase are mixed together in the reaction mixture. The following operations are now performed sequentially.
Step 1: The reaction mixture is heated to a temperature (usually 90-98°C) that assures DNA denaturation.
Step 2 : The mixture is now cooled to a temperature (generally 40-60°C) that permits annealing of the primer to the complementary sequences in the DNA; these sequences are located at the 3 ends of the two strands of the desired segment. This step is called annealing.
Step 3 : The temperature is now so adjusted that the DNA polymerase synthesizes the complementary strands by utilizing 3′-OH of the primers; this reaction is the same as that occurs in vivo during replication of the leading strand of a DNA duplex. The primers are extended towards each other so that the DNA segment lying between the two primers is copied.
The completion of step 3 completes the first cycle of amplification; each cycle may take few (ordinarily 1-3) minutes.
Step 4 : The next cycle amplification is separated the newly synthesized DNA strands from the old DNA strands.
Step 5 : Annealing allows the primers to base pair with both the new and did strands, the total number of strands being twice their original number.
Step 6 : Synthesis of new strands takes place, which doubles the number of copies of the desired DNA segment present at the end of step 1. This completes the second cycle.
Thus at each cycle, both new and old strands anneal to the primers and serve as templates for DNA synthesis. As a result, at the end of each cycle, the number of copies of the desired segment becomes twice the number present at the end of the previous cycle. Thus at the end of n cycles 2n copies of the segment are expected; the real values are quite close to this expectation. The cycle may be repeated.
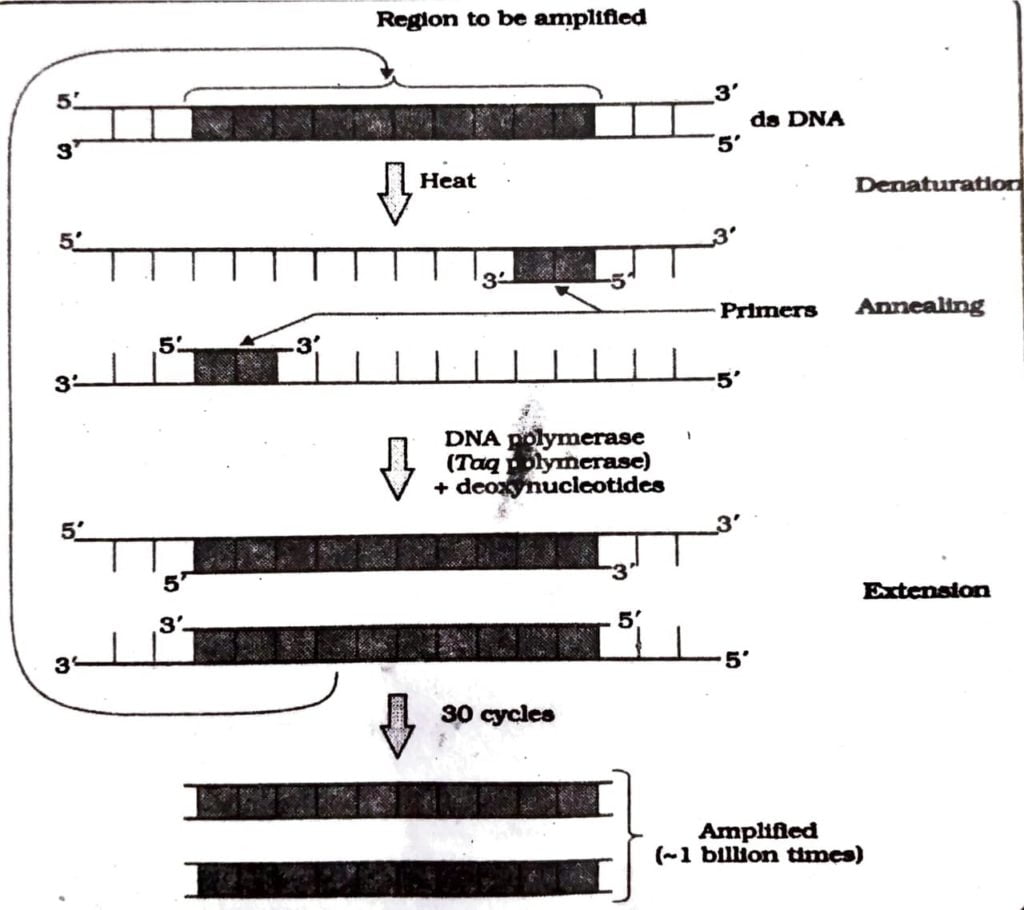
Q.5. Discuss the conceptual development of the principles of genetic engineering.
Ans : Genetic engineering deals with modification of organisms genetic makeup through a highly sophisticate technology that works at ultramicroscopic level to insert or remove nucleotide sequence DNA. The basic features of all organisms is code in the nucleotide sequence in the DNA. Gene is a part of DNA borne by chromosome. The nucleotide sequence of the DNA provides precise information about how certain organism should act and behave. Thus the nature of enzyme amino acid etc. are predetermined in an organism. A type of organisms. may be endowed with particular trait and others may not possess the same depending upon presence or absence particular nucleotide sequence which codes for a specific amino acid chain.
Similarly, certain genetic disorders are caused due to wrong sequence which code for wrong amino acid chain. Replacement of one base pair by another base pair or insertion of one extra pair or deletion of one base pair from DNA results in altered function of the 5. gene. Such minute alternation brings difference between life and death in case of many genetic diseases. Such alternation brings about change in functioning of an organism, its vigour and productivity. How such defective nucleotide sequence could be replaced by a normal one or how desired gene could be implanted in chromosome is the problem of genetic engineering. Genetic engineering can be defined as the science dealing with the correction of nature’s mistake at the level of hereditary unit-the DNA and to improve the quality of organism for specific purposes by implanting desired genes.
It can also be defined as a science dealing with adding to removing genetic units in order to achieve permanent and heritable changes in plants or animals for the benefit or mankind. The opening of a new revolutionary high-skill-technology of genetic engineering was possible due to hara work of Nirenberg, Khorana. Holley and Cohen during 1930-1975 at Massachusetts Institute of Technology and Stanford University of USA. With further work in progress the science of genetics has now become one of the most important applied sciences which will be able to guide the destiny of mankind. The technology has already reached a high level of skill. With further knowledge of the molecular nature of genes and of their capacity to manipulate cells of higher organisms, the possibility of artificial production of new and high yielding varieties of plants, animals and microorganism will further increase. The time is not far when genetic surgery and gene therapy will bring benefit to human society as well.
Genetic engineering so far is centered on to in vitro joining of DNA fragments of different origin with the help of some highly specific enzymes to produce recombinant DNA. Such recombinant DNA is then incorporated into the suitable host. The recombinant DNA technology as it is called has great potentiality in changing our society and in increasing production of food and other items.
The genetic engineering at present deals with a technique called gene cloning. It is the central feature of genetic engineering today. It involves isolation of suitable genes and incorporation of these in suitable hosts to allow their replication.
The technique involves the following steps:
(i) Isolation of required DNA, generally called donor DNA out of a (1) suitable organism (at present most works have been restricted to bacteria and virus) by biochemical technique.
(ii) Breaking of DNA into fragments using restriction enzymes.
(iii) Isolation of suitable bacterial plasmid (extra circular DNA present in many bacteria that are not part of chromosome but can replicate usually and inherit certain traits) and cutting it at a place by restriction enzyme and joining the donor DNA fragment with it with the help of an enzyme called ligase. The plasmid containing parent DNA and donor DNA is called recombinant DNA.
(iv) The recombinant DNA is then inserted within similar bacteria through cloning.
(v) Culturing of eloned bacteria containing fragment of donor DNA.
(vi) Selection of appropriate bacterial colony which has incorporated the sought after DNA fragment. For example, the different segments of DNA inserted in plasmid may cause bacteria to synthesize different substances. It is possible therefore to create bacteria acquiring ability to synthesize any substance like insulin, haemoglobin, interferon etc.
Genetic engineering is comparatively a new discipline of applied science, but within a snore span of time it has shown tremendous progress. Genetic engineering is the basis of biotechnology. Increasing the productivity of biological system, creating newer organisms and elimination of genetic defect from society are the basic aim of genetic engineering now.
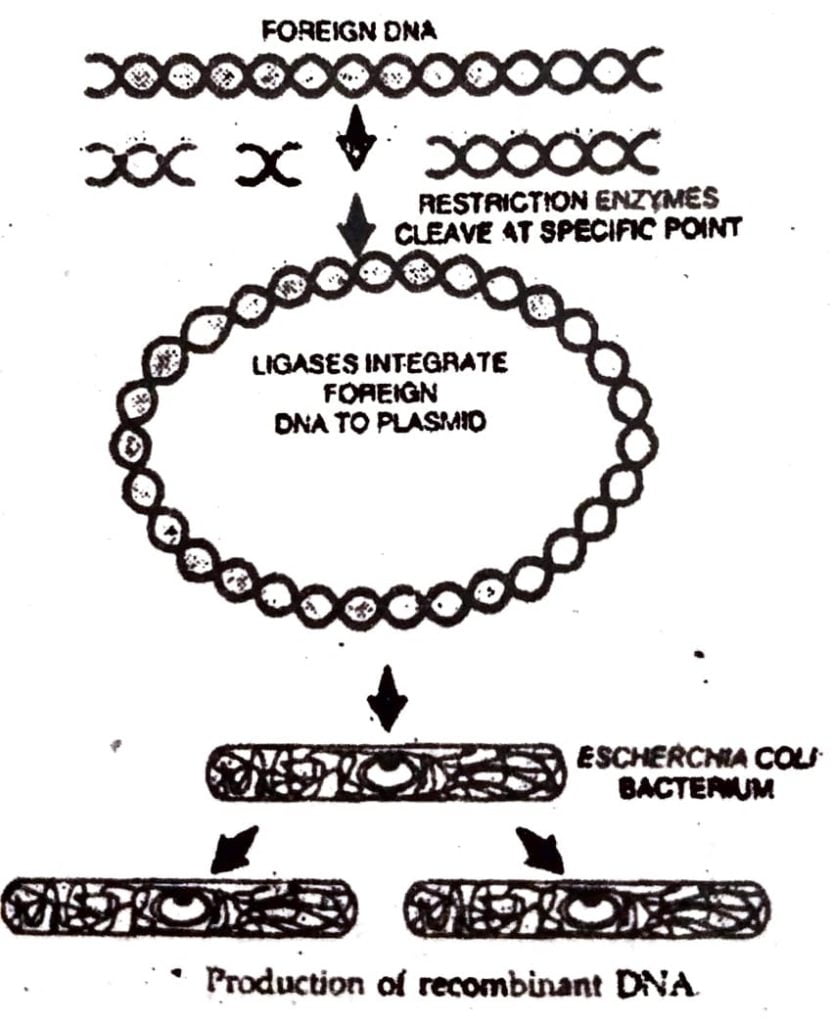
Q.6. What is recombinant DNA technology ? Write a brief account of the process of recombinant DNA technology.
Ans : Recombinant DNA technology includes techniques of cutting DNA into specific fragments using enzymes restriction endonucleases and joining the fragments with the help of enzyme, ligase. Using this technology, the fragments of foreign DNA can be inserted into a vector which may be plasmids or viruses. This technology bypasses the restriction in the gene transfer mechanisms between unrelated organisms. Recombinant DNA technology or genetic engineering involves the following steps.
(i) DNA fragments coding for proteins of interest are synthesized 6. chemically or isolated from an organism.
(ii) These DNA fragments are inserted in a restriction endonuclease cleavage site of the vector that does not inactivate any gene required for vector’s maintenance.
(iii) The recombinant DNA molecules are now introduced into a host to replicate.
(iv) Recipient host cells that have acquired the recombinant DNA, are selected. Selection pressure is applied to enrich bacteria with a selectable market.
(v) Desired clones are then characterised to ensure that they maintain true copies of the DNA segment that was originally cloned.
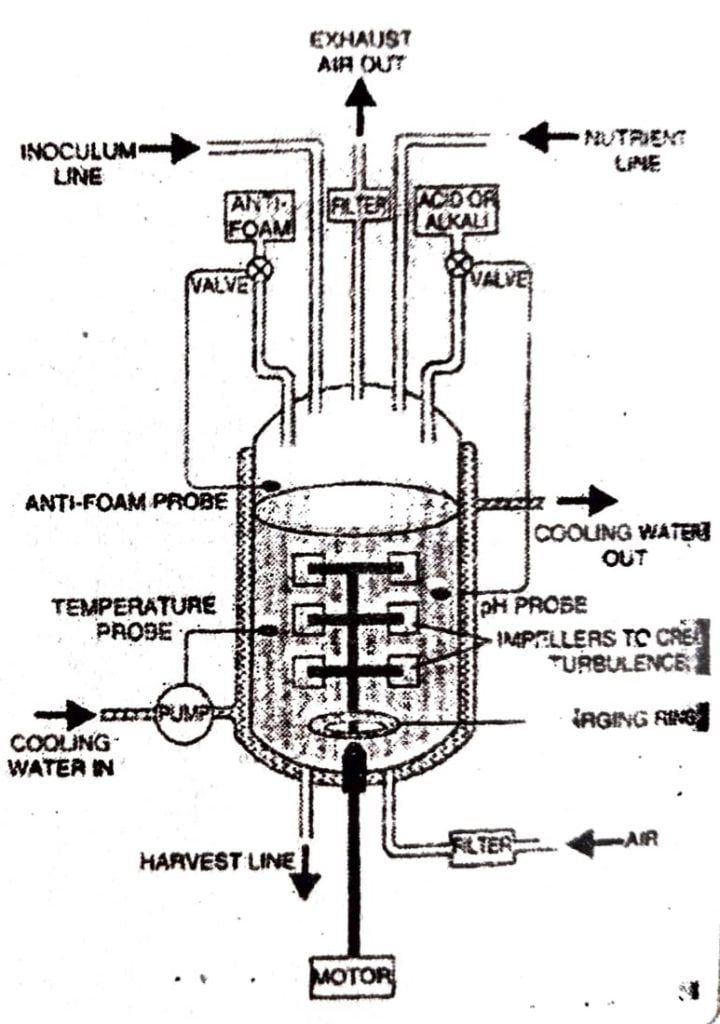
Q.7. What is bioreactor ? Draw a labelled diagram of stirred bioreactor and explain its functioning.
Ans : Structure: The basic design of a stirred-tank fermenter is shown below: It consists of a large stainless steel vessel with a capacity of upto 500,000 dm’ around which there is a jacket of circulatory water used to control the temperature within the fermenter. There is also an agitator, comprising of a series of flat blades, which can be rotated with the help of a motor. This ensures the thoroughly mixing of the contents so that nutrients come in close with the micro-organisms. The agitator also 7. sparged prevents settling out of the cells at the bottom.
Fermenter also has adequate arrangement for aeration, temperature and pH control. For proper aeration, air can be forced in at the bottom of the tank through a porous ring, called sparger, by the process called sparging, while there is an outlet to remove air and waste gases at the top of the tank.
The top of the tank also a number of inlet tubes called ports, through which materials can be introduced or withdrawn e.g.
– Inoculation port for introducing initial inoculum.
– Nutrient port for introducting more nutrients.
– Antifoam for port introducting antifoaming agents.
– pH port for introducing acid or alkali to maintain optimal pH.
At the base of the tank, there is a harvest line to extract culture medium and microbial products. To regularly detect the pH and temperature changes, tank is fitted with ceruan probes.
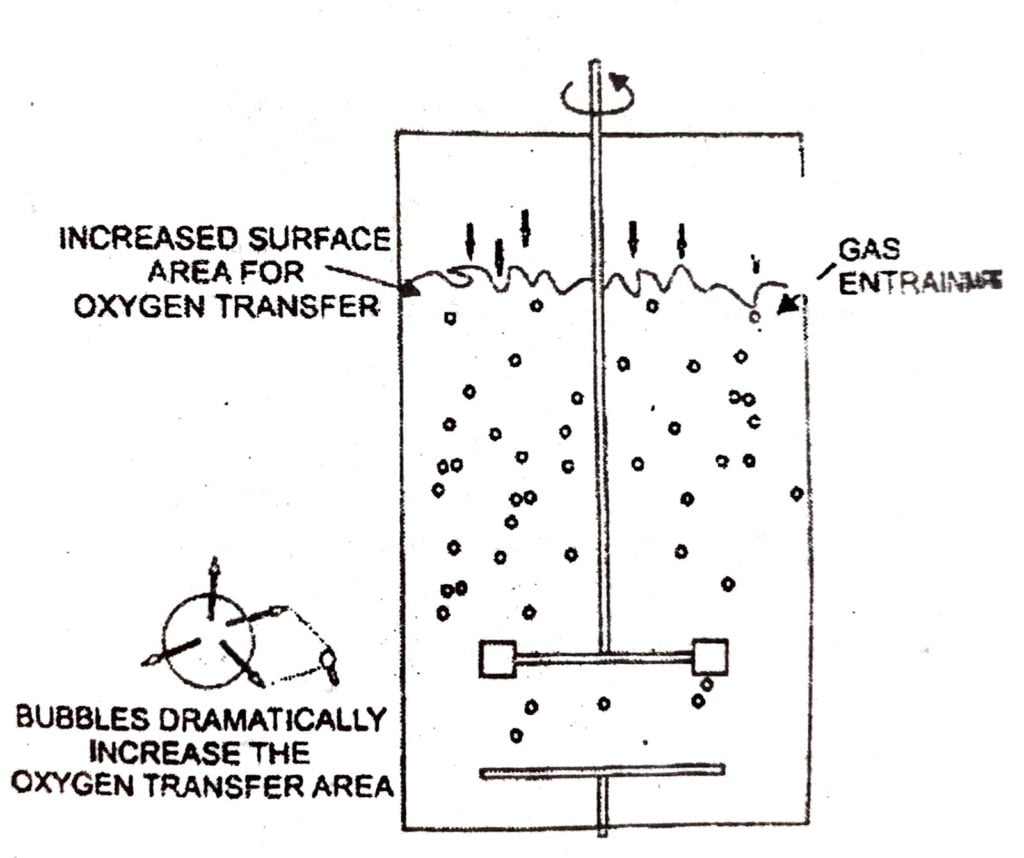
Significance : The bioreactor is a well tried and tested design for large scale production of micro organisms under aseptic condition and controlled environment for a number of days. Small scale fermenters of 10-100 lits Capacity are use in research laboratories. It is also provided with cortrols for the monitoring of physical, chemical and biological parameters that affect the growth of cells.

Hi, I’m Dev Kirtonia, Founder & CEO of Dev Library. A website that provides all SCERT, NCERT 3 to 12, and BA, B.com, B.Sc, and Computer Science with Post Graduate Notes & Suggestions, Novel, eBooks, Biography, Quotes, Study Materials, and more.


 Buy Now
Buy Now
Hii, i wan’t to say that please make the answers in brief.. The answers are too long